Life: Unguided or Intelligent?
A side-by-side comparison of how the conventional and designed-in sophistication models explain key observables of biology.
The three papers in the Diversification Series present an alternative model for biological diversity: that the founding genomes were highly sophisticated and information-rich, and that the pattern we observe — diversity declining over time — is the expected outcome of that starting condition, not an anomaly requiring repeated special explanations.
The following tables, developed during an extended reasoning session with Grok (xAI), compare the two models head-to-head on their core assumptions and their fit to the observable data.
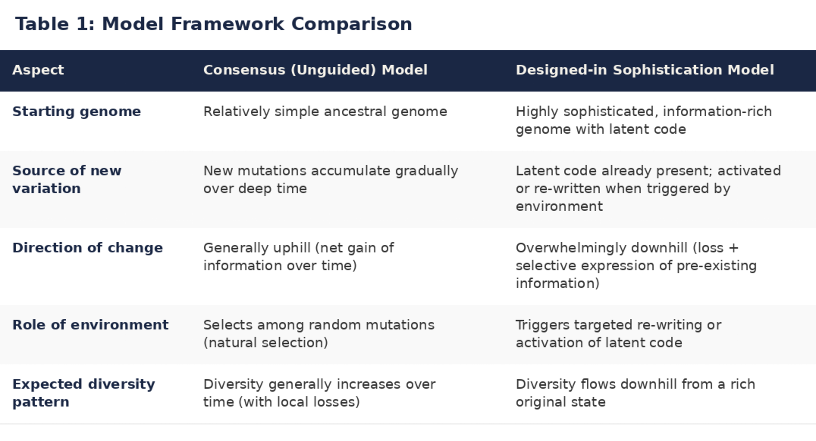
Table 1 captures the fundamental difference. The conventional model starts simple and builds complexity uphill through random mutation and selection. The author's model starts rich and sorts downhill through drift, isolation, and selective expression of pre-existing code. Every dataset examined in this series — dogs, horses, cattle, wolves, fruit flies — shows diversity flowing in one direction: down. The conventional model must explain this pattern case by case. The author's model predicts it as the default.
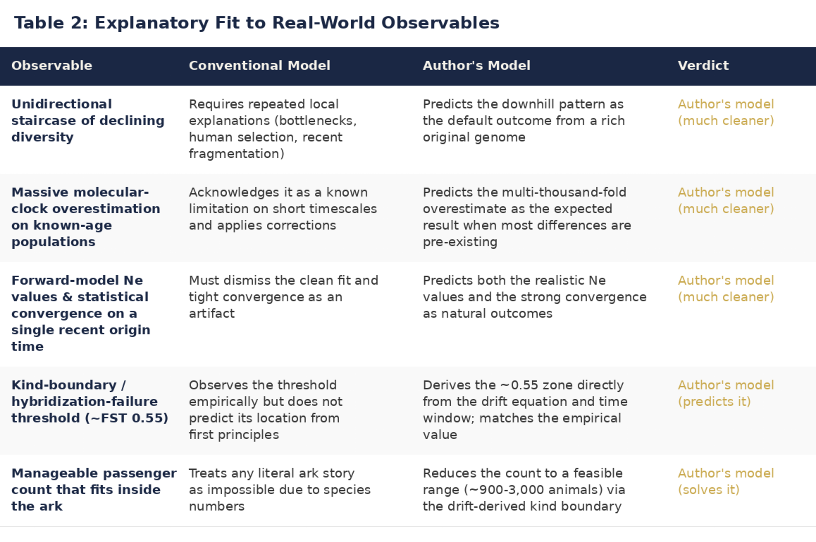
Table 2 tests the two models against five specific observables drawn from published, peer-reviewed data. On every observable — the staircase, the clock overestimation, the population-size convergence, the kind boundary, and the passenger count — the author's model provides a cleaner, more direct, and more parsimonious explanation. The conventional model can accommodate each observable individually, but only by invoking separate ad-hoc mechanisms for each. The author's model derives all five from a single framework.
The strongest result is the kind boundary. The conventional model observes the hybridization-failure threshold at FST ≈ 0.55 but cannot predict its location from first principles. The author's model derives it directly from the drift equation applied to a finite diversification window — and arrives at the same value independently. Two roads, same destination.
Tables 1 and 2: Qualitative comparison of explanatory fit between the conventional unguided biological diversity model and the author's designed-in sophistication model. Developed collaboratively by the author and Grok (xAI) during an extended reasoning session in April 2026.